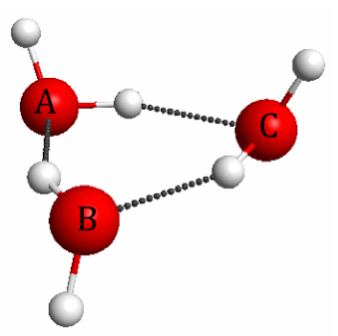
Figure 4.17. M06/6-31G(d) optimized geometry of the water trimer.
Click on the picture for an interactive versionFrom
Molecular Modeling Basics CRC Press, 2010
In a previous post I showed how molecular electrostatic potentials can be used to illustrate
polarization due intermolecular interactions. Polarization has an interesting and important consequence called the many-body effect, which occurs in systems with many relatively strong interactions such as the water trimer (Figure 4.17).
The many body effect is due to the fact that polarization of the charge density of, say, water molecule A due to the A–B hydrogen bond increases the strength of the A–C hydrogen bond, which lowers the energy further. As a result the binding energy of the water trimer (i.e. its energy relative to that of three free water molecules) is lower than the sum of the three hydrogen bond strengths.
The A–B hydrogen bond strength can be gotten by (1) deleting water C and computing the energy of the A-B dimer (don't re-optimize), and (2) comparing this energy to that of two free water molecules. And similarly for the A–C and B–C dimers. The coordinates can be extracted from the Jmol interactive figure as shown in
a previous post, and
this post shows how to perform the calculations with GAMESS.
The increased hydrogen bond strengths are also reflected in the hydrogen bond lengths, which are 0.09Å shorter in the trimer compared to the dimer. You can measure the distances between atoms in the Jmol interactive figure by double clicking on the atoms, and water dimer geometry can be found in
a previous post.